BEBPA Blog
Volume 2, Issue 8
Characterization of Critical Reagents for HCP ELISA Development by Orthogonal Methods
By Pia Paarmann, Ph. D., Head of 2D Analytics, BioGenes GmbH
The development of host cell protein enzyme-linked immunosorbent assays (HCP ELISAs) requires comprehensive analytical characterization of the critical ELISA reagents including the HCP mock material and anti-HCP antibodies by different orthogonal methods (as described in USP 1132[1]).
The process-specific mock material is a critical reagent which serves many different purposes. It is the antigen source material for immunization and affinity purification and is used as the ELISA standard. A well-prepared mock sample successfully represents most process-specific HCPs. To check for this requirement, the similarity of the HCP populations between the mock sample and an early-stage sample from the product production process (such as a classical harvest sample) can be assessed prior to immunization. 2D DIGE is a powerful visual method for an analysis of overall mock and process sample characteristics and a comparison of HCP spot patterns. The method shows the individual distribution of proteins over a wide molecular weight range and pI range. A high degree of similarity indicates successful representation of the process-specific HCPs.
An orthogonal assessment of protein identities by LC-MS analysis (as detailed in USP 1132.1[2]) in mock and process samples further increases the level of in-depth characterization. Using LC-MS as an analytical tool in the early identification of persisting HCPs in the final drug substance sample enables a targeted search of potentially problematic HCPs in the respective mock sample preparation.
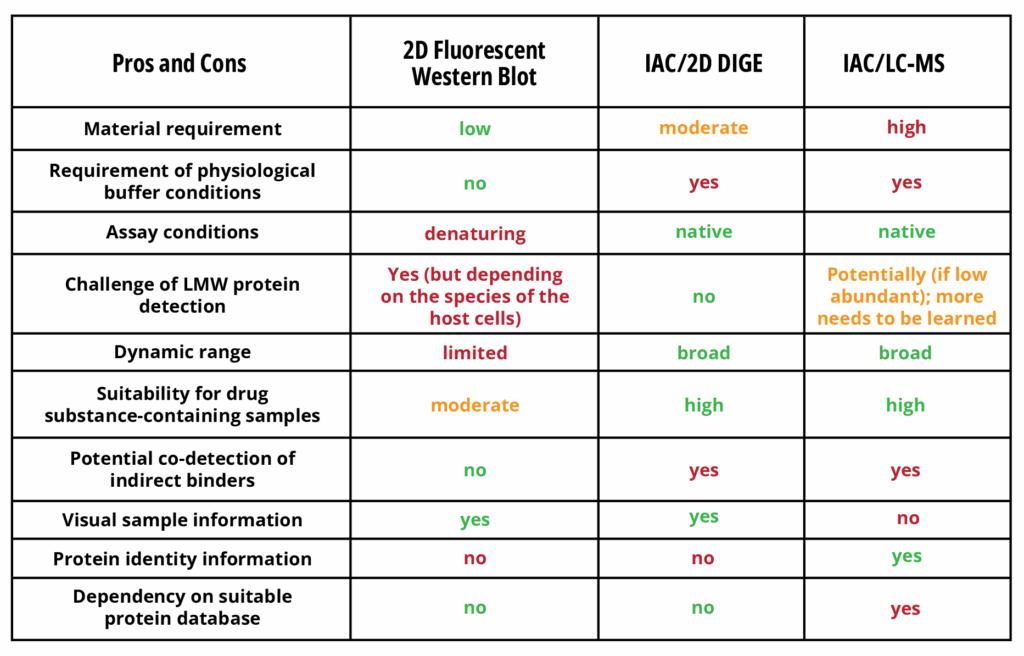
The final capture antibody is another critical ELISA reagent. Checking for reasonable dilutional linearity, high sensitivity, HCP specificity and a reliable reflection of HCP log reduction over the downstream process is essential during ELISA development. HCP coverage determination is a further crucial part of capture antibody characterization, as the detection of a broad spectrum of proteins is a scientific and regulatory requirement. There are various analytical methods for assessing coverage, all of which have their individual strengths and weaknesses (Table 1).
A traditional method to look directly at capture antibody coverage performance is to run a 2D Western blot. Making use of sensitive fluorescent minimal labeling or post-electrophoretic fluorescent total protein staining for visualization enables the detection of antigen and immunostaining signal patterns directly from the same Western blot membrane. This has largely improved the accuracy of signal correlations. High-resolution technical equipment for 2D electrophoresis has further enhanced the significance of 2D images, especially regarding the visualization of low molecular weight (LMW) proteins.
However, the detection of LMW proteins by 2D Western blot methods can be challenging, especially for mammalian HCP samples, due to the natural technical restrictions that are present with the use of this method:
- fewer epitopes of smaller proteins which are susceptible to denaturing assay conditions
- limited amount of sample load (especially when analyzing drug-containing process samples)
- potentially lower immunogenicity of smaller proteins
- potentially lower abundance of smaller proteins
Meanwhile, the overall expectations on percentage coverage values from 2D Western blot analyses have changed, now revealing more realistic numbers, carefully taking into account the impact of technical improvements such as the enhanced resolution of LMW proteins and more accurate spot matching, as well as the host cell line-specific properties of samples. Individual, case-by-case considerations and a comprehensive analytical data set are necessary. Immunoaffinity Chromatography (IAC), using an immobilized capture antibody for extraction of successfully bound HCPs, is now widely applied as an orthogonal approach for coverage determination. The read-out after IAC can be performed by 2D DIGE, thereby comparing the HCP spot patterns of the input sample and its corresponding IAC eluate fraction. The IAC method benefits from native assay conditions resembling the chemical environment of the ELISA and has a much broader dynamic range of detection as it enables the enrichment of low abundant proteins while also distinguishing HCPs from drug substances.
Significantly different percentage coverage values may thus be obtained depending on whether a 2D fluorescent Western blot or IAC/2D DIGE is applied.
IAC can clearly compensate for some of the technical limitations of a 2D Western blot analysis. However, the elution of high-affinity binders from the capture column can be challenging, while the risk of co-eluting indirect binders cannot be fully excluded. For IAC, the physiological buffer conditions of the sample to be analyzed are a requirement. In case of inclusion bodies, this requirement can be an insolvable technical problem.
Lastly, individual spots or spot groups which are clearly detected in the 2D Western blot assay may not be detected using IAC/2D DIGE and vice versa. Orthogonal methods are valuable in that they can complement each other.
A further level of information and complexity is currently offered by using LC-MS as a read-out method after IAC.
LC-MS is highly valued in the identification of proteins (including already known potential high-risk HCPs[3]) in different input samples and corresponding IAC eluate fractions, thereby supporting an answer to the question of whether an HCP antibody is fit for purpose or not.
Careful process optimization and risk assessment can be carried out accordingly. The development of single HCP ELISAs for the quantitative monitoring of potential high-risk proteins may be a requirement[4],[5].
Integrating the knowledge and percentage coverage numbers obtained by different orthogonal methods represents a stimulating and important challenge. For example, the total spot numbers in 2D DIGE or 2D Western blot analyses depend on the operating procedures in their respective laboratories, as well as on the software used for data analysis and the individual interpretation of biological/chemical phenomena. The numbers and hits derived from LC-MS analyses can also vary significantly. Hence, there remains much to learn regarding the diversity of HCP projects.
Each analytical method presents its own technical challenges, strengths, and limitations. Placing the coverage values into the right context and adding orthogonal pieces of information to the puzzle of HCP impurity analyses will help to provide safe biological therapeutics to patients.
Table 1: Orthogonal methods for HCP coverage determination[1]
[1] U.S. Pharmacopeia, <1132> Residual host cell protein measurement in biopharmaceuticals, USP 39–NF 34, 2016.
[2] U.S. Pharmacopeia, <1132.1> Residual host cell protein measurement in biopharmaceuticals by liquid chromatography–mass spectrometry, 2025.
[3] Jones M, Palackal N, Wang F, Gaza-Bulseco G, Hurkmans K, Zhao Y, Chitikila C, Clavier S, Liu S, Menesale E, Schonenbach NS, Sharma S, Valax P, Waerner T, Zhang L, Connolly T. “High-risk” host cell proteins (HCPs): A multi-company collaborative view. Biotechnol Bioeng. 2021 Aug;118(8):2870-2885. doi: 10.1002/bit.27808. Epub 2021 May 31. PMID: 33930190.
[4] de Zafra CL, Quarmby V, Francissen K, Vanderlaan M, Zhu-Shimoni J. Host cell proteins in biotechnology-derived products: A risk assessment framework. Biotechnol Bioeng. 2015 Nov;112(11):2284-91. doi: 10.1002/bit.25647. Epub 2015 Jun 16. PMID: 26010760
[5] Wang F, Richardson D, Mueller H-M, Gaza-Bulseco G, Liu X, Liu F, et al. Host-cell protein risk management and control during bioprocess development: a consolidated biotech industry review, part 1&2. BioProcess Int. 2018

About The Author: Pia Paarmann
Dr. Pia Paarmann is heading the 2D analytics department at BioGenes, based in Berlin, Germany. Together with her team, Pia uses two-dimensional gel electrophoresis techniques (such as 2D-DIGE and 2D-DIBE) for comprehensive HCP sample and antibody reagent characterization. Pia has gathered experience from numerous industry customer’s projects and has closely looked into the advantages and limitations of different analytical methods.
Dr. Paarmann holds a doctoral degree in biochemistry with a focus on growth factor signal transduction and endocytosis from the “Freie Universität” of Berlin.
